Générer des animations NDVI
Produits utilisés : ndvi_anomaly
Mots-clés : données utilisées ; anomalie ndvi, agriculture
Aperçu
Ce bloc-notes montre comment générer une animation pour visualiser le NDVI dans l’espace, le NDVI moyen dans le temps et les anomalies du NDVI. Il génère également une animation similaire pour les précipitations mensuelles et les anomalies pluviométriques.
Les animations peuvent aider à visualiser les changements de tendances au fil du temps. La visualisation des valeurs moyennes et des anomalies signifie que les aspects saisonniers et interannuels du temps peuvent être évalués. En complétant cela avec des images spatiales, on améliore l’interprétabilité des données d’observation de la Terre.
Description
Le flux de travail pour générer une animation progresse selon les étapes suivantes :
Sélectionnez une zone d’intérêt.
Charger le produit NDVI moyen et anomalies DE Africa.
Masque pour terres cultivées.
Générer une animation NDVI.
Générer une animation de précipitations. ***
Commencer
Pour exécuter cette analyse, exécutez toutes les cellules du bloc-notes, en commençant par la cellule « Charger les packages et les applications ».
Charger des paquets
[1]:
%matplotlib inline
import os
import datacube
import datetime as dt
import xarray as xr
import geopandas as gpd
from odc.geo.geom import Geometry
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from IPython.display import HTML
import matplotlib.dates as mdates
from deafrica_tools.plotting import display_map
from deafrica_tools.areaofinterest import define_area
Connectez-vous au datacube.
[2]:
dc = datacube.Datacube(app='NDVI-animation')
Paramètres d’analyse
La zone par défaut est les terres cultivées à Madagascar.
La cellule suivante définit les paramètres qui définissent la zone d’intérêt et la durée de l’analyse. Les paramètres sont
« lat » : la latitude centrale à analyser (par exemple « 10,338 »).
« lon » : la longitude centrale à analyser (par exemple « -1,055 »).
« buffer » : le nombre de degrés carrés à charger autour de la latitude et de la longitude centrales. Pour des temps de chargement raisonnables, définissez cette valeur sur « 0,1 » ou moins.
Sélectionnez l’emplacement
Pour définir la zone d’intérêt, deux méthodes sont disponibles :
En spécifiant la latitude, la longitude et la zone tampon. Cette méthode nécessite que vous saisissiez la latitude centrale, la longitude centrale et la valeur de la zone tampon en degrés carrés autour du point central que vous souhaitez analyser. Par exemple, « lat = 10,338 », « lon = -1,055 » et « buffer = 0,1 » sélectionneront une zone avec un rayon de 0,1 degré carré autour du point avec les coordonnées (10,338, -1,055).
Vous pouvez également fournir des valeurs de tampon distinctes pour la latitude et la longitude d’une zone rectangulaire. Par exemple, « lat = 10.338 », « lon = -1.055 » et « lat_buffer = 0.1 » et « lon_buffer = 0.08 » sélectionneront une zone rectangulaire s’étendant sur 0,1 degré au nord et au sud et 0,08 degré à l’est et à l’ouest à partir du point « (10.338, -1.055) ».
Pour des temps de chargement raisonnables, définissez la mémoire tampon sur « 0,1 » ou moins.
By uploading a polygon as a
GeoJSON or Esri Shapefile. If you choose this option, you will need to upload the geojson or ESRI shapefile into the Sandbox using Upload Files button in the top left corner of the Jupyter Notebook interface. ESRI shapefiles must be uploaded with all the related files
in the top left corner of the Jupyter Notebook interface. ESRI shapefiles must be uploaded with all the related files (.cpg, .dbf, .shp, .shx). Once uploaded, you can use the shapefile or geojson to define the area of interest. Remember to update the code to call the file you have uploaded.
Pour utiliser l’une de ces méthodes, vous pouvez décommenter la ligne de code concernée et commenter l’autre. Pour commenter une ligne, ajoutez le symbole "#" avant le code que vous souhaitez commenter. Par défaut, la première option qui définit l’emplacement à l’aide de la latitude, de la longitude et du tampon est utilisée.
Si vous exécutez le bloc-notes pour la première fois, conservez les paramètres par défaut ci-dessous. Cela permettra de démontrer le fonctionnement de l’analyse et de fournir des résultats significatifs.
[3]:
# # Define the area of interest for the analysis
year = "2022"
# Method 1: Specify the latitude, longitude, and buffer
aoi = define_area(lat=-17.71, lon=48.21, buffer=0.05)
# Method 2: Use a polygon as a GeoJSON or Esri Shapefile.
# aoi = define_area(vector_path='aoi.shp')
#Create a geopolygon and geodataframe of the area of interest
geopolygon = Geometry(aoi["features"][0]["geometry"], crs="epsg:4326")
geopolygon_gdf = gpd.GeoDataFrame(geometry=[geopolygon], crs=geopolygon.crs)
# Get the latitude and longitude range of the geopolygon
lat_range = (geopolygon_gdf.total_bounds[1], geopolygon_gdf.total_bounds[3])
lon_range = (geopolygon_gdf.total_bounds[0], geopolygon_gdf.total_bounds[2])
display_map(lon_range, lat_range)
[3]:
Nous définirons la date de début et de fin comme le début et la fin de notre année d’intérêt.
[4]:
start_date = dt.datetime.strptime(year, "%Y").replace(month=1, day=1)
end_date = dt.datetime.strptime(year, "%Y").replace(month=12, day=31)
Ci-dessous, les paramètres d’analyse sont utilisés pour construire une requête qui charge le produit NDVI moyen et les anomalies de DE Africa.
[5]:
# Construct the data cube query
query = {
'x': lon_range,
'y': lat_range,
'time': (start_date, end_date),
'output_crs': 'EPSG:6933',
'resolution': (-30, 30)
}
ds = dc.load(product="ndvi_anomaly", **query)
ds
[5]:
<xarray.Dataset> Size: 14MB
Dimensions: (time: 12, y: 406, x: 323)
Coordinates:
* time (time) datetime64[ns] 96B 2022-01-16T11:59:59.999999 .....
* y (y) float64 3kB -2.218e+06 -2.218e+06 ... -2.231e+06
* x (x) float64 3kB 4.647e+06 4.647e+06 ... 4.656e+06
spatial_ref int32 4B 6933
Data variables:
ndvi_mean (time, y, x) float32 6MB 0.4654 0.4267 0.3652 ... nan nan
ndvi_std_anomaly (time, y, x) float32 6MB -0.1621 0.2036 0.3302 ... nan nan
clear_count (time, y, x) int8 2MB 4 4 4 4 4 4 4 4 ... 0 0 0 0 0 0 0 0
Attributes:
crs: epsg:6933
grid_mapping: spatial_refCharger le masque des terres cultivées
Il s’agit d’une étape facultative pour masquer les données NDVI sur les terres cultivées. Dans ce cas, nous souhaitons uniquement inspecter les schémas de végétation des cultures, nous utilisons donc le masque pour le faire.
[6]:
query = {
'x': lon_range,
'y': lat_range,
'time': '2019',
'output_crs': 'EPSG:6933',
'resolution': (-30, 30),
'measurements': 'mask'
}
cm = dc.load(product='crop_mask', **query).squeeze()
cm.mask.where(cm.mask<255).plot()
[6]:
<matplotlib.collections.QuadMesh at 0x7f2e6f4b87d0>
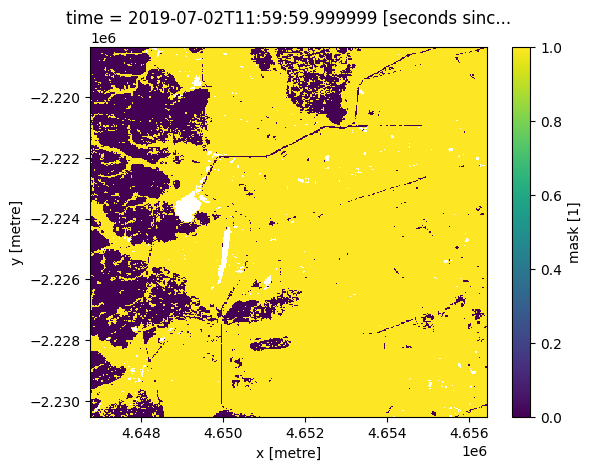
Masquer le NDVI pour les terres cultivées
Nous masquons les données NDVI à l’étendue des terres cultivées indiquée ci-dessus.
[7]:
ds = ds.where(cm.mask ==1)
Préparer les données d’animation
Générer une animation de précipitations. ***
NDVI moyen avec attributs spatiaux.
NDVI moyen au fil du temps, agrégé sous forme de moyenne dans l’espace.
Anomalies NDVI pour chaque mois, agrégées sous forme de moyenne dans l’espace.
[8]:
ds_mean = ds.ndvi_mean
ds_mean_1D = ds_mean.mean(dim=['x', 'y'], skipna=True)
ndvi_anom = ds.ndvi_std_anomaly.mean(dim=['x', 'y']).to_dataframe().drop(['spatial_ref'], axis=1)
Exécuter l’animation
Pour créer l’animation, la cellule ci-dessous suit quelques étapes clés :
Définir la disposition des trois axes et leur taille.
Définir les paramètres des axes (titres, etc.)
Commencez chaque axe avec la première image de l’animation (janvier).
Définissez une fonction de mise à jour qui ajoute chaque mois séquentiellement à chacun des trois graphiques.
Exécutez l’animation.
Notez que l’animation s’affiche dans sa propre fenêtre. Le bouton en bas à droite permet de télécharger l’animation.
[9]:
# create a figure and axes
fig = plt.figure()
fig.set_figheight(5)
fig.set_figwidth(10)
ax1 = plt.subplot2grid(shape=(2, 2), loc=(0, 1), colspan=2)
ax2 = plt.subplot2grid(shape=(2, 2), loc=(0, 0), rowspan=2)
ax3 = plt.subplot2grid(shape=(2, 2), loc=(1, 1), colspan=2)
# setup starting plot
cax = ds.ndvi_mean[0,:,:].plot(
add_colorbar=True,
cmap='RdYlGn',
vmin=-1, vmax=1,
cbar_kwargs={
'extend':'neither'
},
ax=ax2
)
ax1.set_ylim([0, 1])
ax1.xaxis.set_major_formatter(mdates.DateFormatter("%b"))
ax1.set_title("Mean NDVI")
ax1.set_xlabel("Date")
ax1.set_ylabel("NDVI")
ax3.set_ylim([-4, 4])
ax3.xaxis_date()
ax3.xaxis.set_major_formatter(mdates.DateFormatter("%b"))
ax3.set_title("NDVI Anomaly")
ax3.set_xlabel("Date")
ax3.set_ylabel("Std Deviations from monthly mean")
ax3.axhline(0, color='black', linestyle='--')
x = ds_mean_1D.time
y = ds_mean_1D
x2 = ndvi_anom.index
y2 = [0] * ndvi_anom.ndvi_std_anomaly
bars = ax3.bar(x2, y2, align='center', width=20,color=(ndvi_anom.ndvi_std_anomaly > 0).map({True: 'g', False: 'brown'}))
line, = ax1.plot(x, y, marker='*',
color='blue')
def update(num, x, y, x2, y2, bars, line):
y2[num] = ndvi_anom.ndvi_std_anomaly[num]
bars[num].set_height(y2[num])
cax.set_array(ds.ndvi_mean[(num),:,:].values.flatten())
ax2.set_title("Time = " + str(ds.ndvi_mean.coords['time'].values[(num)])[:12])
line.set_data(x[:num+1], y[:num+1])
return line,
plt.tight_layout()
ani = animation.FuncAnimation(fig, update, len(x), fargs=[x, y, x2, y2, bars, line],
interval=400, blit=False, repeat=True)
#ani.save('ndvi_anim.gif')
plt.close()
HTML(ani.to_html5_video())
/tmp/ipykernel_7580/754916817.py:47: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
y2[num] = ndvi_anom.ndvi_std_anomaly[num]
/tmp/ipykernel_7580/754916817.py:47: FutureWarning: Series.__setitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To set a value by position, use `ser.iloc[pos] = value`
y2[num] = ndvi_anom.ndvi_std_anomaly[num]
/tmp/ipykernel_7580/754916817.py:48: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
bars[num].set_height(y2[num])
[9]:
Créer une animation NDVI avec les précipitations
L’animation ci-dessus résume le NDVI des terres cultivées dans l’espace et le temps, sur une période donnée. Nous pouvons ajouter des informations associées à cette visualisation, telles que les précipitations. Les prochaines étapes montrent comment charger les données de précipitations et calculer les anomalies pour le traçage.
Paramètres d’anomalies de précipitations
Nous devons définir certains paramètres pour calculer l’anomalie des précipitations. La période « time_m » est une période de référence tandis que « time_x » est l’année d’intérêt.
[10]:
# Set the range of dates for the climatology, this will be the reference period (m) for the anomaly calculation.
# Standard practice is to use a 30 year period, so we've used 1981 to 2011 in this example.
time_m = ('2000', '2020')
# time period for monthly anomaly (x)
time_x = ('2022')
# CHIRPS has a spatial resolution of ~5x5 km
resolution = (-5000, 5000)
#size of dask chunks
dask_chunks = dict(x=500,y=500)
query_m = {
'x': lon_range,
'y': lat_range,
'time': time_m,
'output_crs': 'EPSG:6933',
'resolution': (-5000, 5000)
}
query_x = {
'x': lon_range,
'y': lat_range,
'time': time_x,
'output_crs': 'EPSG:6933',
'resolution': (-5000, 5000)
}
Charger les données de précipitations
[11]:
ds_rf_m = dc.load(product='rainfall_chirps_monthly', **query_m)
ds_rf_x = dc.load(product='rainfall_chirps_monthly', **query_x)
Ci-dessous, nous calculons la moyenne mensuelle des précipitations et l’anomalie.
[12]:
# monthly means
climatology_mean = ds_rf_m.groupby('time.month').mean('time').compute()
#calculate monthly std dev
climatology_std = ds_rf_m.groupby('time.month').std('time').compute()
stand_anomalies = xr.apply_ufunc(
lambda x, m, s: (x - m) / s,
ds_rf_x.groupby("time.month"),
climatology_mean,
climatology_std,
output_dtypes=[ds_rf_x.rainfall.dtype],
dask="allowed"
).compute()
Préparer les données d’animation
Quant à l’animation NDVI ci-dessus, nous devons préparer des trames de données pour l’animation des précipitations.
[13]:
spatial_mean_rain = ds_rf_x.rainfall.mean(['x','y'])
spatial_mean_rain.time
spatial_mean_anoms = stand_anomalies.rainfall.mean(['x','y'], skipna=True).sel(time=slice('2021-12-31', end_date)).to_dataframe().drop(
['spatial_ref', 'month'], axis=1).fillna(0)
Exécutez l’animation
Nous suivons les étapes répertoriées pour l’animation NDVI ci-dessus pour produire l’animation des précipitations.
[14]:
# create a figure and axes
fig = plt.figure()
fig.set_figheight(5)
fig.set_figwidth(10)
ax1 = plt.subplot2grid(shape=(2, 2), loc=(0, 1), colspan=2)
ax2 = plt.subplot2grid(shape=(2, 2), loc=(0, 0), rowspan=2)
ax3 = plt.subplot2grid(shape=(2, 2), loc=(1, 1), colspan=2)
cax = ds.ndvi_mean[0,:,:].plot(
add_colorbar=True,
cmap='RdYlGn',
vmin=-1, vmax=1,
cbar_kwargs={
'extend':'neither'
},
ax=ax2
)
ax1.set_title("Monthly Total Rainfall")
ax1.set_xlabel("Date")
ax1.xaxis.set_major_formatter(mdates.DateFormatter("%b"))
ax1.set_ylabel("Monthly Rainfall (mm)")
ax3.clear()
ax3.set_ylim([-4, 4])
ax3.xaxis_date()
ax3.xaxis.set_major_formatter(mdates.DateFormatter("%b"))
ax3.set_title("Rainfall Anomaly")
ax3.set_xlabel("Date")
ax3.set_ylabel("Std Deviations from monthly mean")
ax3.axhline(0, color='black', linestyle='--')
def update(num, x, y, line):
y2.iloc[num] = spatial_mean_anoms.rainfall.iloc[num]
bars[num].set_height(y2.iloc[num])
cax.set_array(ds.ndvi_mean[(num),:,:].values.flatten())
ax2.set_title("Time = " + str(ds.ndvi_mean.coords['time'].values[(num)])[:12])
line.set_data(x[:num+1], y[:num+1])
return line,
x = spatial_mean_rain.time
y = spatial_mean_rain
x2 = spatial_mean_anoms.index
y2 = [0] * spatial_mean_anoms.rainfall
bars = ax3.bar(x2, y2, align='center', width=20,color=(spatial_mean_anoms.rainfall > 0).map({True: 'b', False: 'r'}))
line, = ax1.plot(x, y, marker='*',
color='blue')
plt.tight_layout()
ani = animation.FuncAnimation(fig, update, len(x), fargs=[x, y, line],
interval=400, blit=True)
plt.close()
HTML(ani.to_html5_video())
[14]:
Tirer des conclusions
Les animations nous permettent d’inspecter l’évolution de la végétation au cours d’une année et, en montrant les anomalies, de comparer l’année d’intérêt à une base de référence historique.
Ce type d’analyse peut aider à évaluer la production agricole au cours d’une année donnée et peut alimenter les systèmes d’alerte précoce et les prévisions sur la production agricole et la sécurité alimentaire.
Informations Complémentaires
Licence Le code de ce notebook est sous licence Apache, version 2.0 <https://www.apache.org/licenses/LICENSE-2.0>`__.
Les données de Digital Earth Africa sont sous licence Creative Commons Attribution 4.0 <https://creativecommons.org/licenses/by/4.0/>.
Contact Si vous avez besoin d’aide, veuillez poster une question sur le canal Slack DE Africa <https://digitalearthafrica.slack.com/> ou sur le GIS Stack Exchange <https://gis.stackexchange.com/questions/ask?tags=open-data-cube> en utilisant la balise open-data-cube (vous pouvez consulter les questions posées précédemment `ici <https://gis.stackexchange.com/questions/tagged/open-data-
Si vous souhaitez signaler un problème avec ce notebook, vous pouvez en déposer un sur Github.
Version de Datacube compatible
[15]:
print(datacube.__version__)
1.8.20
Dernier test :
[16]:
from datetime import datetime
datetime.today().strftime('%Y-%m-%d')
[16]:
'2025-01-16'